
Argomenti
- Common omics data formats
Obiettivi conoscitivi
Al termine di questa attività dovresti essere in grado di:
- Descrivere le caratteristiche e gli scopi dei principali formati di dati omici (es. FASTQ, GTF/GFF e BED)
Standard formats for omics data
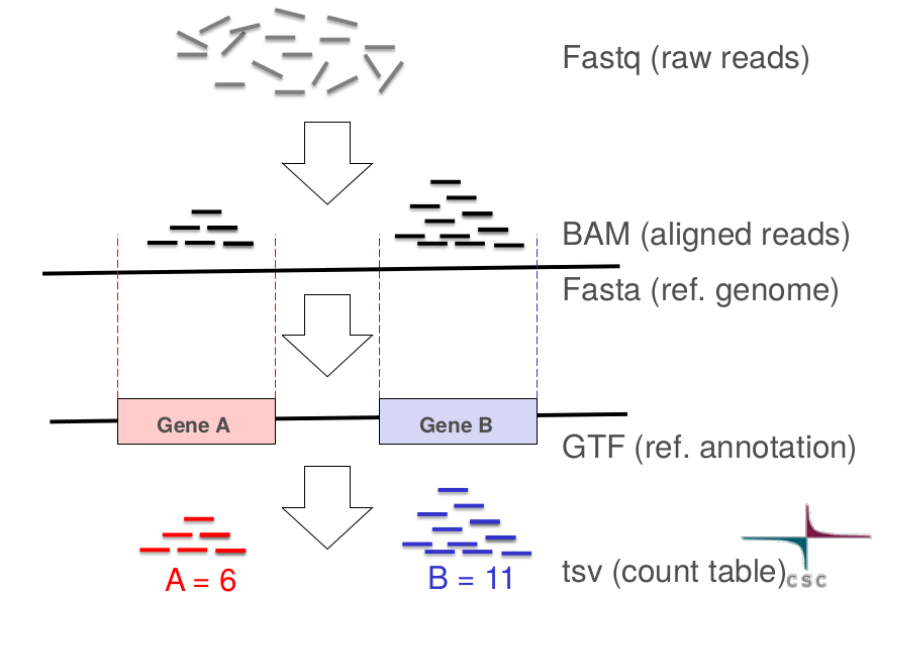
In genomics and transcriptomics, standard data formats are essential for ensuring interoperability between tools, reproducibility of analyses, and ease of data sharing. Many types of omics data are stored in simple text-based formats, which are widely supported and human-readable. For example, nucleotide sequences are often stored in FASTA or FASTQ files, genomic intervals in BED, and gene annotations in specialized formats such as GFF/GTF files. Using standardized formats allows researchers to efficiently analyze, visualize, and integrate diverse datasets.
❓ Challenge question: How many sequences would you expect to find in a FASTA file containing the human genome? Why?
❓ Challenge question: Knowing the size of the human genome and assuming ~1 byte per character, what is the expected size of a FASTA file containing the entire genome?
A small FASTQ file to play with
This is a microbiome sample from Jacques et al. 2021.
Download gunzipped FASTQ file (]500kb)
❓ Challenge question:
- Download and unzip the file
- Inspect the uncompressed file from a bash terminal (hint: use head, tail)
- How many reads does the file contain? (hint: use the bash command wc -l)
❓ super-challenging question:
- Download and install the FASTQC software on your PC (see the instructions after clicking Download Now)
- Launch the FastQC application
- Load the FASTQ file you previously downloaded (select File > Open. You can then select the files you want to analyse.)
- Run the quality control analysis
- Evaluate results (see here)
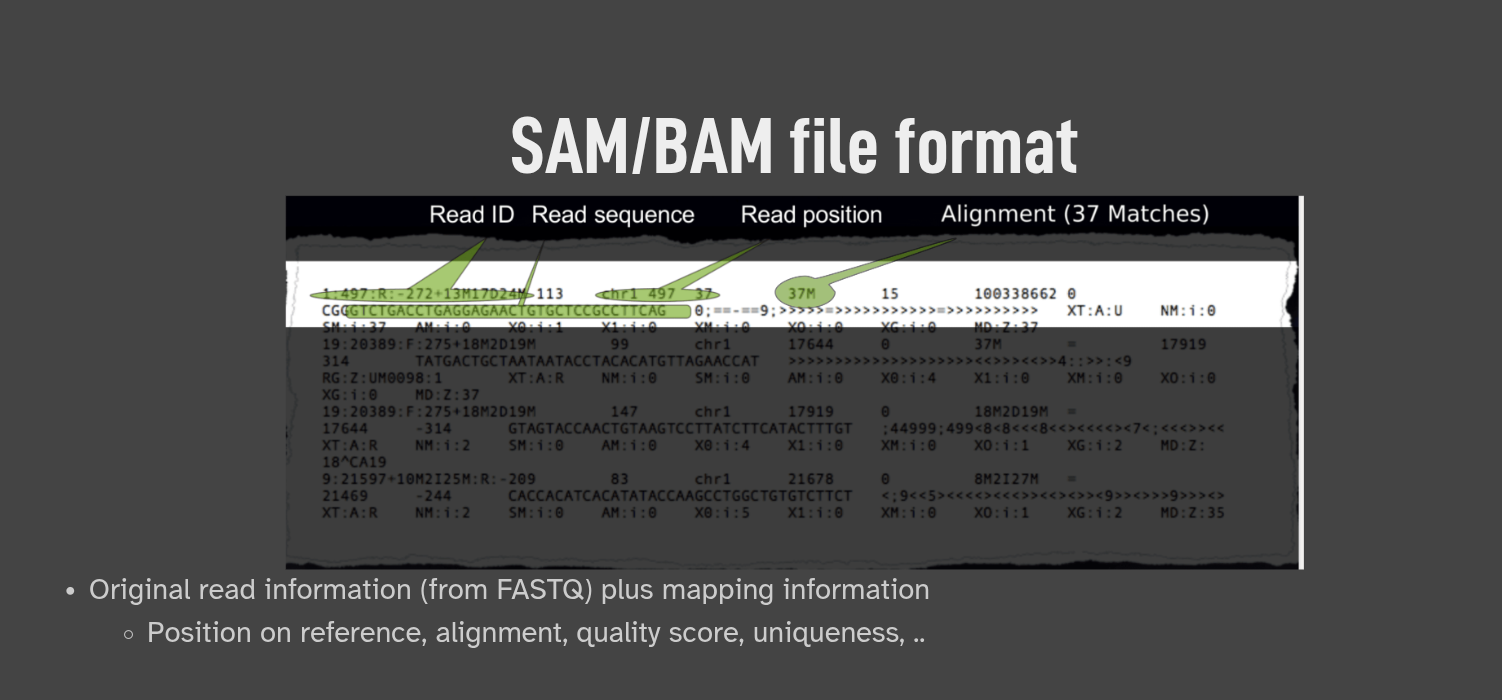

(see the instructions after clicking Download Now)