
Topics
Exploring the UCSC Genome Browser and Associated Tools
Learning objectives
By the end of this activity, you should be able to:
- Visualize a genomic locus in the UCSC Genome Browser and identify key genomic features (e.g., genes, exons, regulatory elements)
- Customize the genomic view by selecting desired annotation tracks (e.g. variation, genes annotated by GENCODE) and deselecting unwanted ones (e.g.
- Map an unknown DNA sequence to a reference genome using a genome browser and interpret its genomic context (e.g., gene location, nearby features)
- Load and visualize a custom track of genomic intervals in the UCSC Genome Browser
- Assemble a custom BED file for selected genomic features of interest
UCSC and the Birth of the Genome Browser
The University of California, Santa Cruz (UCSC) Genome Browser was developed by graduate student Jim Kent, and its development is intertwined with the final and crucial steps of the Human Genome Project. The first version of this graphical tool, designed to navigate the newly released draft of the human genome, was published on July 7, 2000.

A still from a short video on the history of the UCSC Genome Browser and its role in the Human Genome Project, available here. </div>
## Navigating Genomic Information: the UCSC Genome Browser
Access the UCSC Genome Browser data portal here. </div>
## Core Tools at UCSC: - **Genome Browser**: The main visualization tool that displays any portion of a genome at any scale with aligned annotation tracks showing genes, regulatory elements, conservation, variants, and other genomic features.
- **BLAT (BLAST-Like Alignment Tool)**: A rapid sequence alignment tool developed by Jim Kent for finding sequence matches in genomes. Faster than BLAST for closely related sequences and useful for locating mRNA/EST alignments.
- **LiftOver**: A tool for converting genomic coordinates between different genome assemblies (e.g., hg19 to hg38). Requires chain files that map regions between assemblies.
- **In-Silico PCR**: A tool for virtually testing PCR primer pairs against a genome to verify specificity and predict amplicon locations.
- **Table Browser**: A web interface for querying, filtering, and downloading data from the underlying MySQL databases. Allows intersection of data tables and export in multiple formats. See below for more related terms, or our documentation.
### **Genome Browser** See an overview of the Genome Browser interface, highlighting its layout, navigation, and key functional elements at [this link](https://github.com/maghetta/teaching/blob/2b66786ddff304090013a51e7272b18e9514d661/navigare_i_genomi/2.%20A%20spasso%20nel%20tuo%20genoma__selected_pages.pdf).
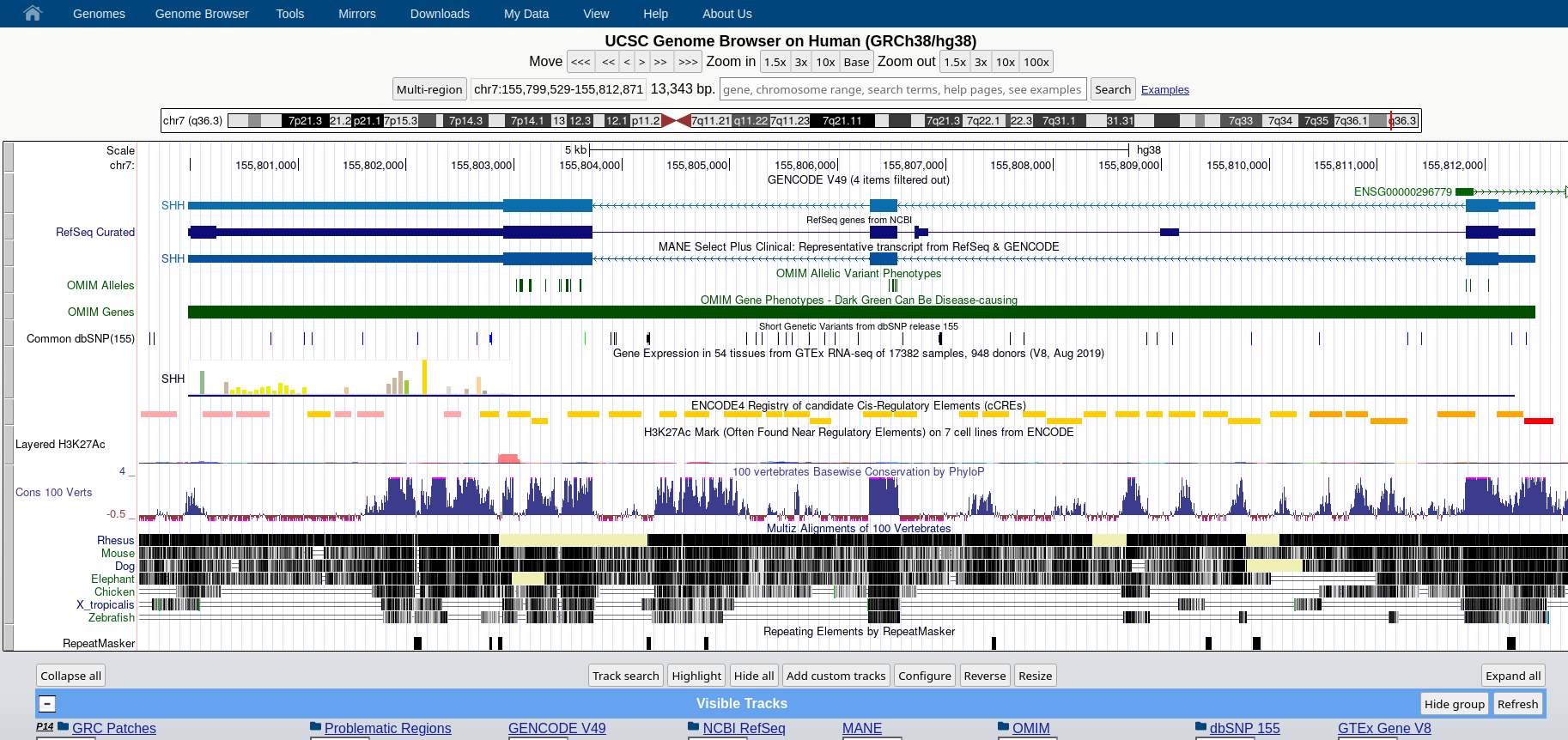
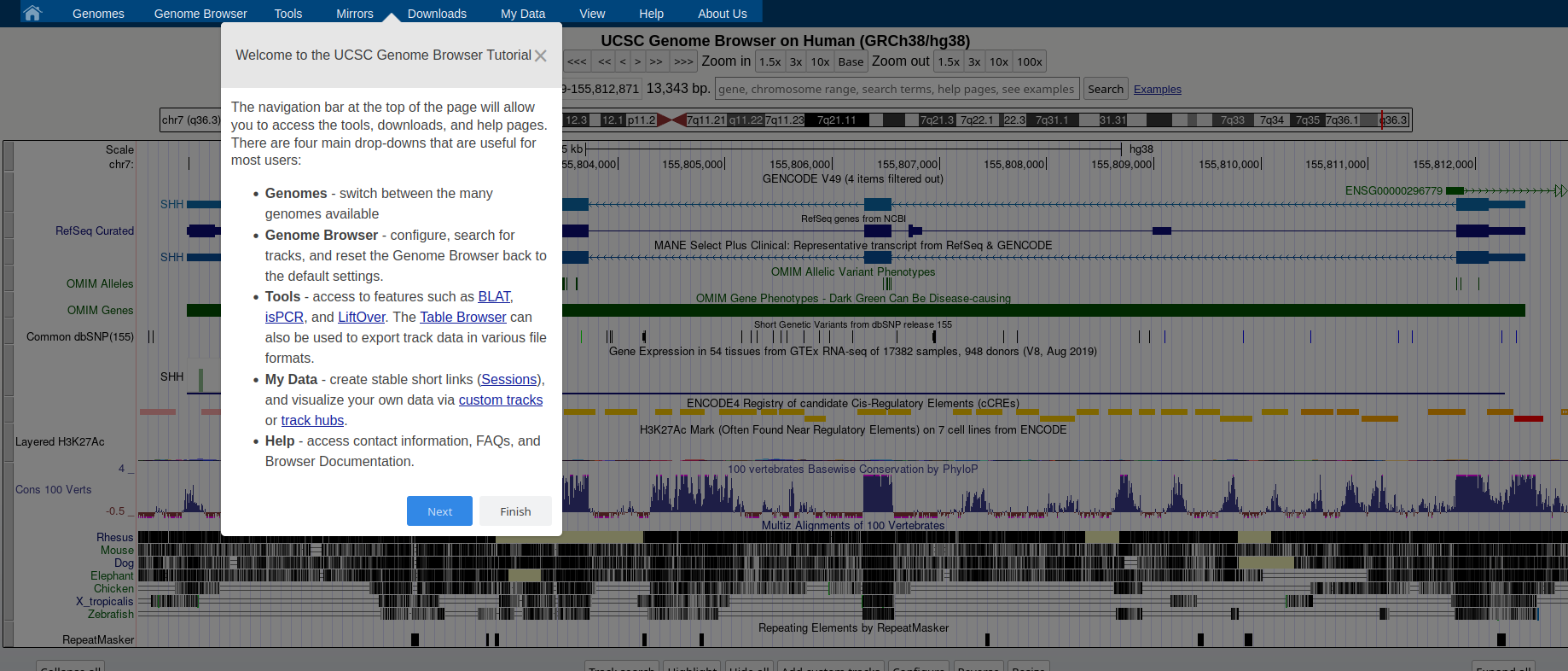
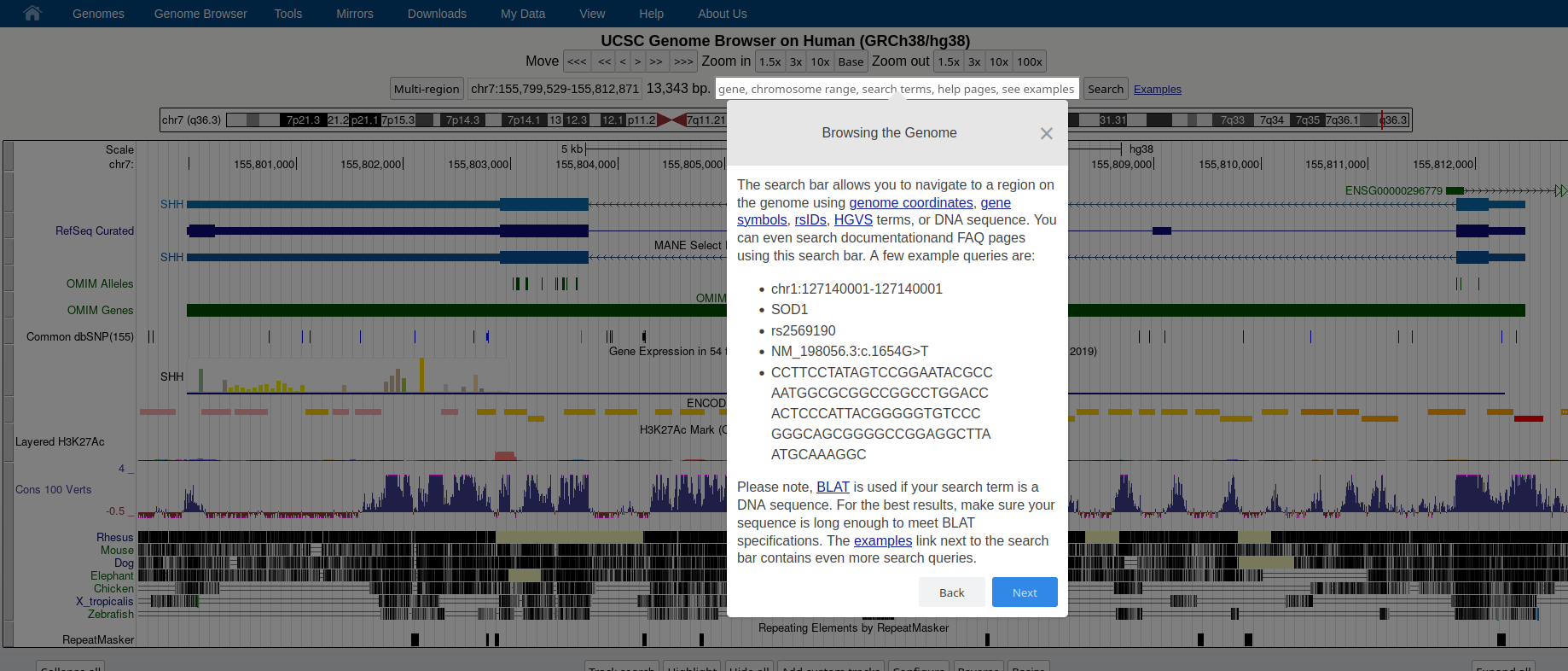
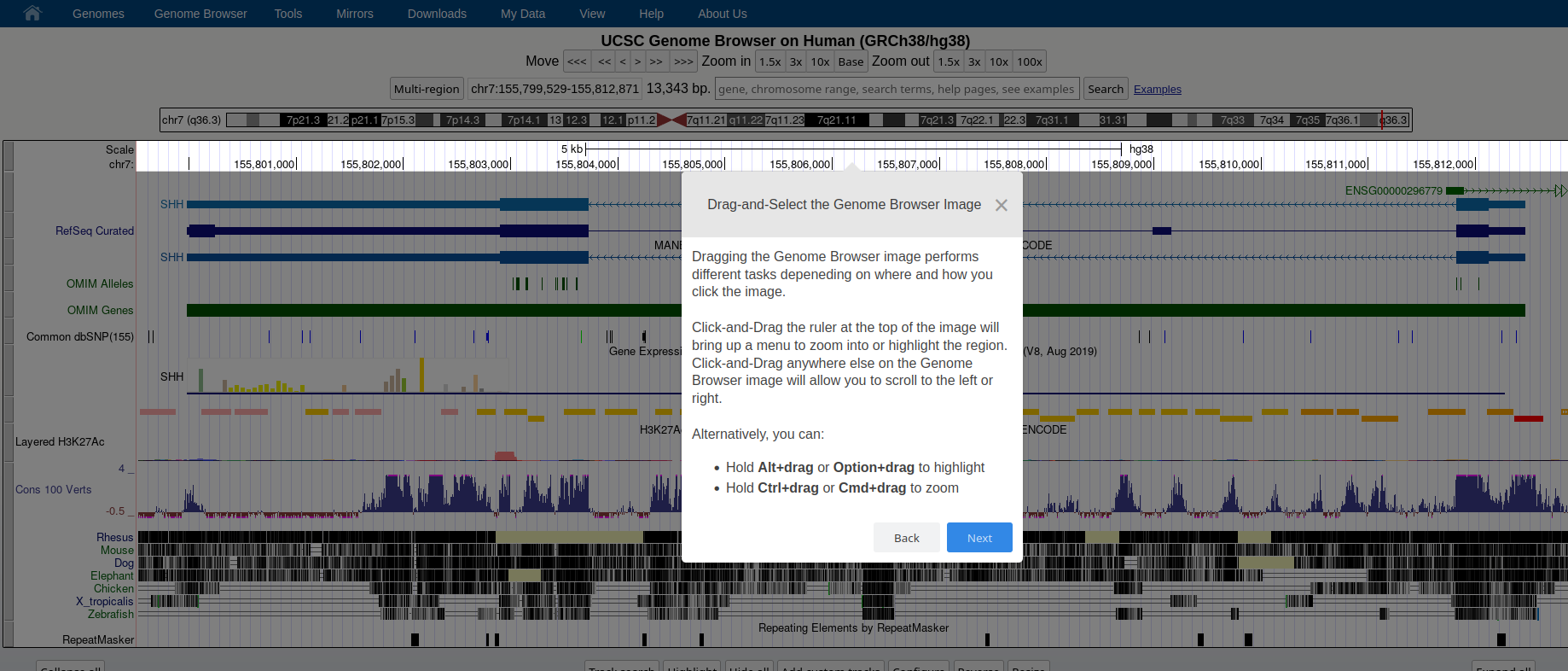
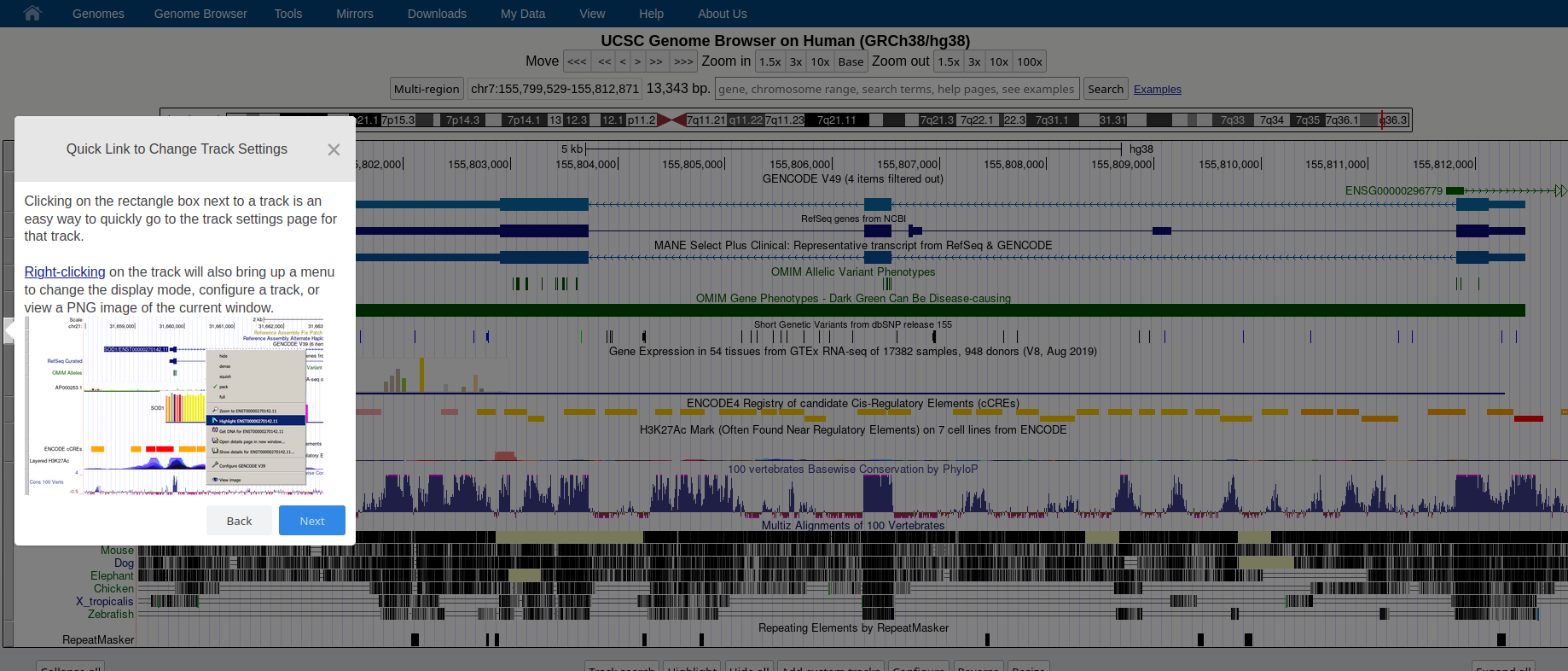
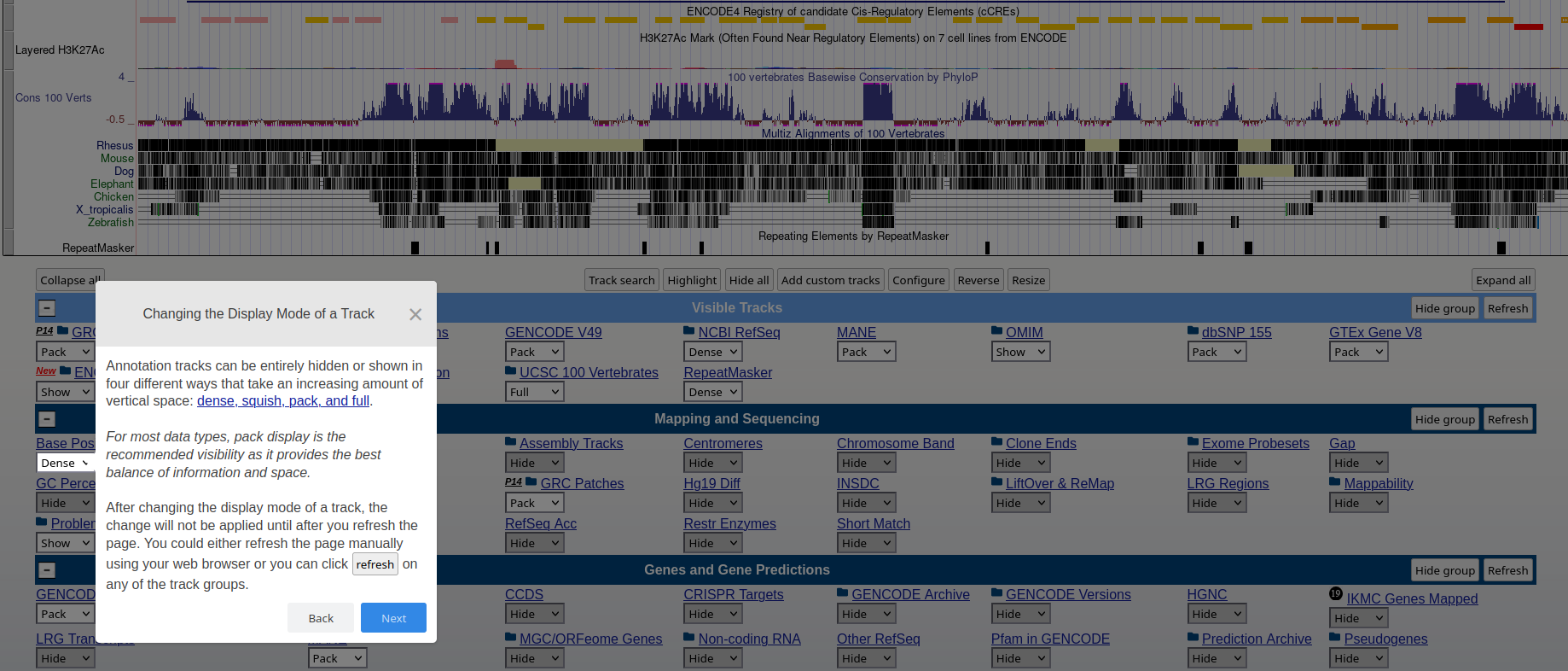
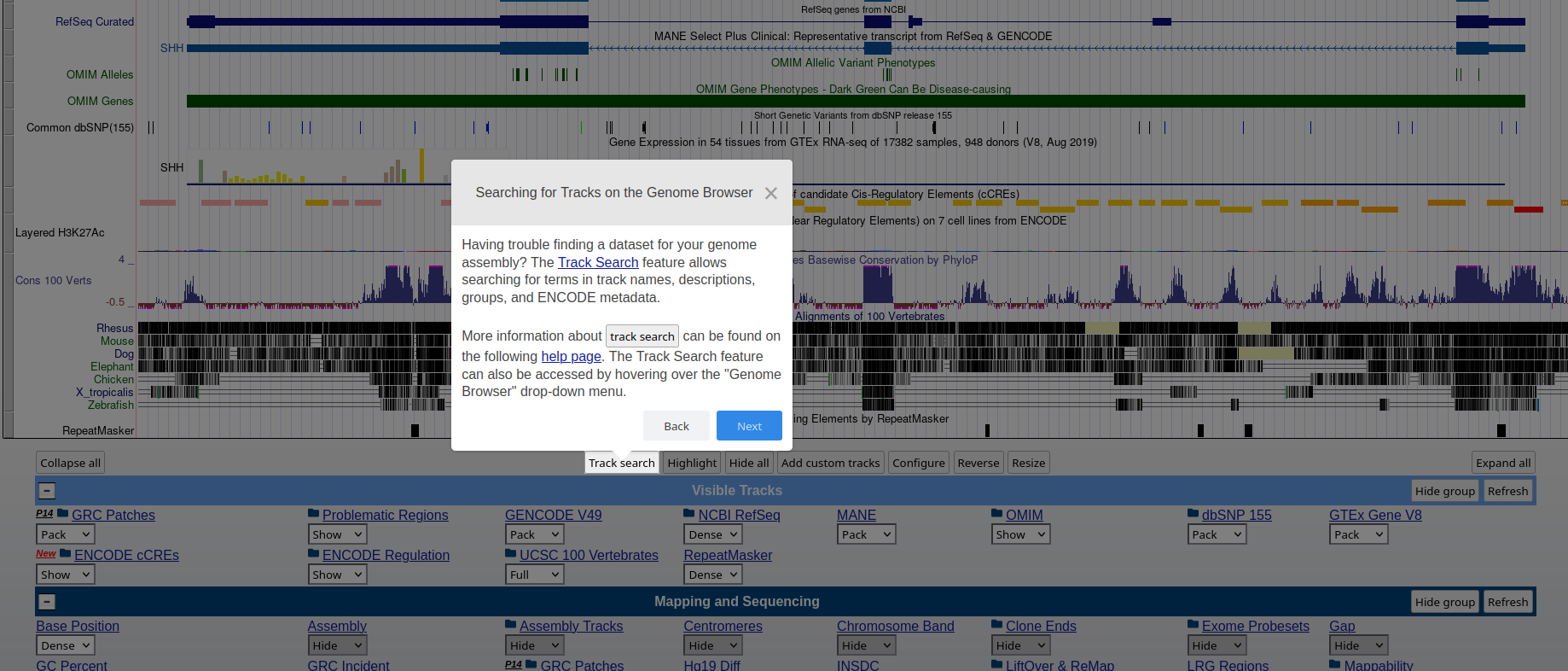
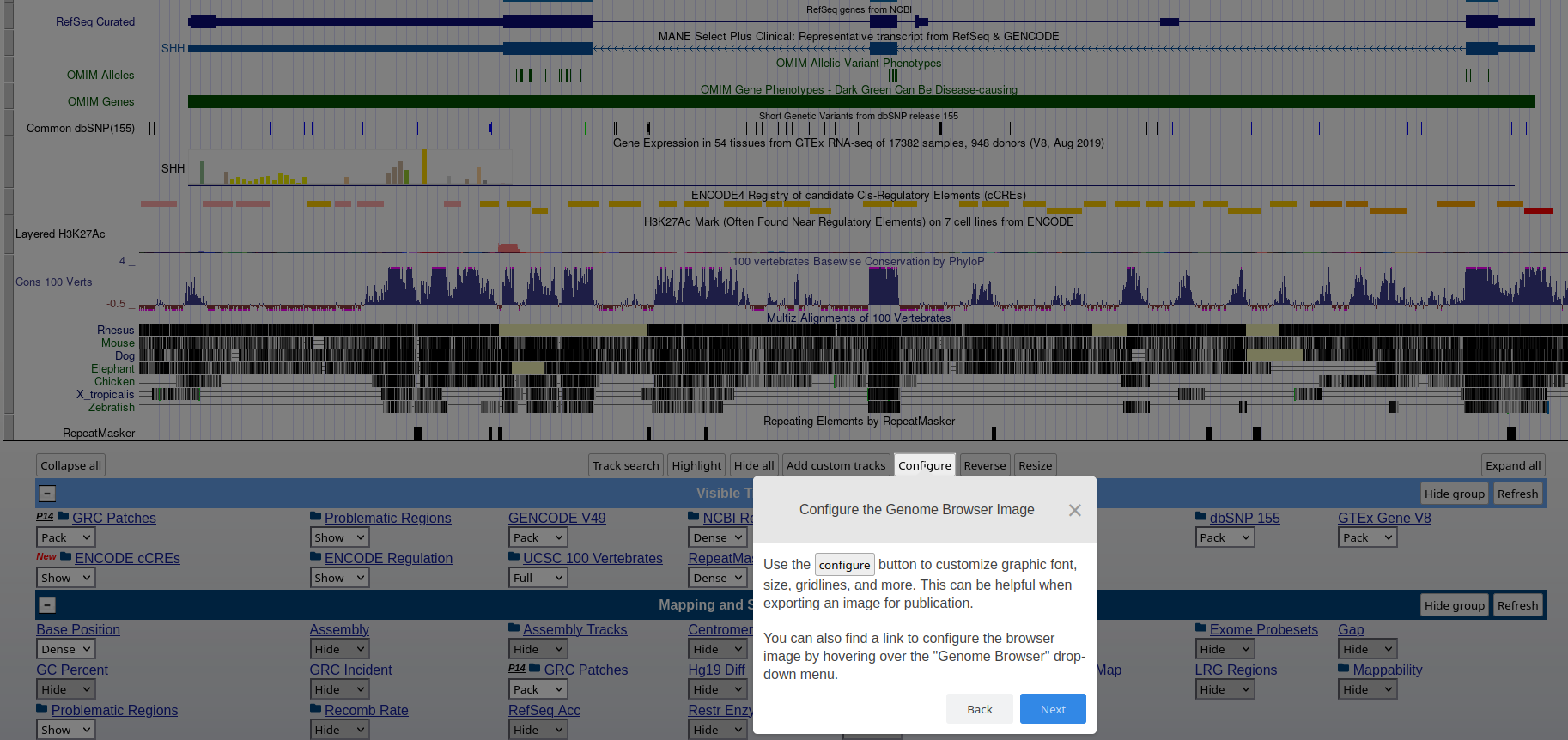
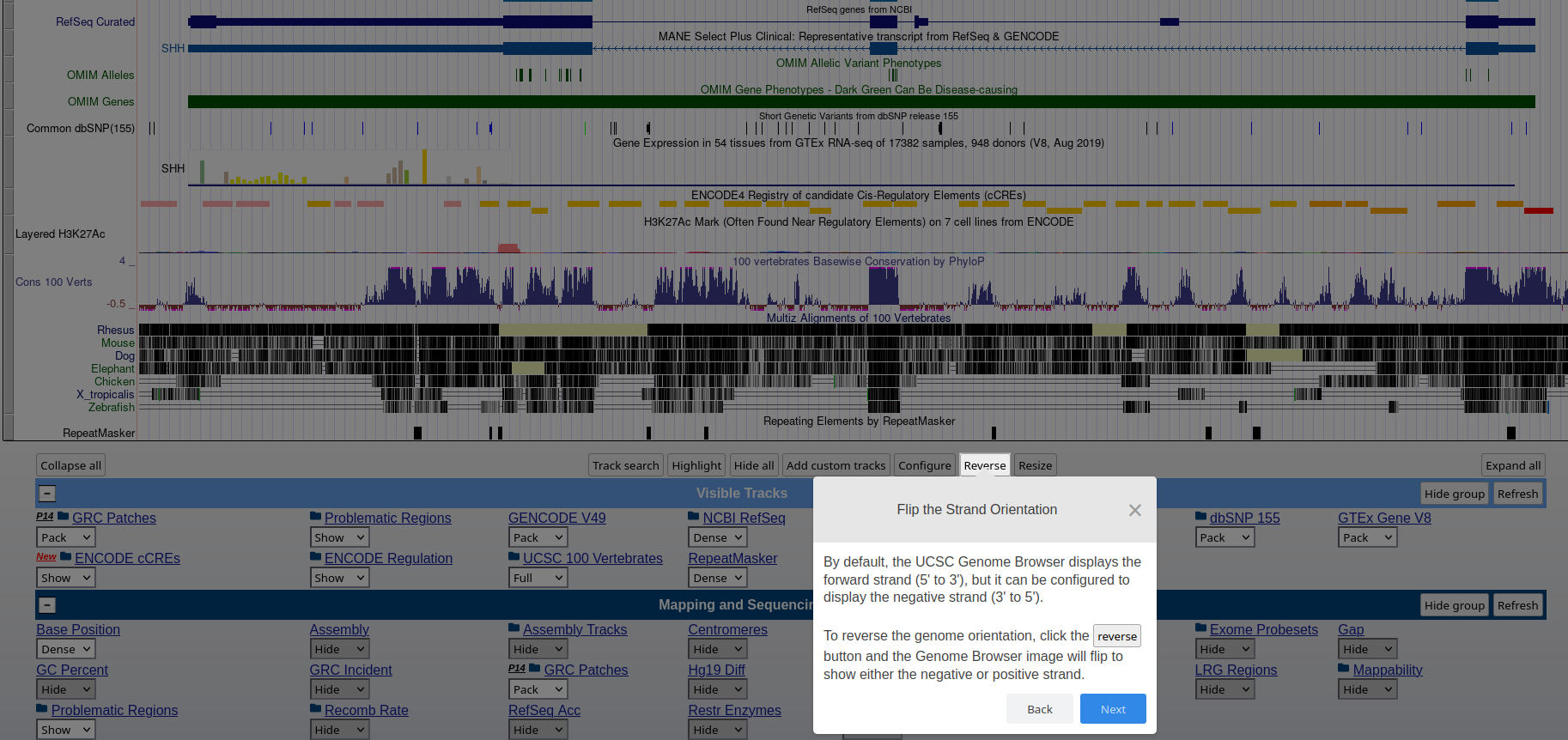
## How to: find sequence matches to the genome (BLAT) and create and upload a custom BED file to the UCSC Genome Browser
See here
the following activities:
- *Attività: dove mappa questa sequenza sul genoma?!*
- *Attività: crea un file BED per una lista di geni di interesse*
## Check PCR primers by using the in-silico PCR tool available at the UCSC genome browser Try the behaviour of the tool (top menu --> tools --> in-silico PCR) by using the Human genome assembly hg38 and the folloeing primers:
- Forward primer: CATCCGCAAAGACCTGTACG
- Reverse primer: CCTGCTTGCTGATCCACATC
**References**
- **UCSC Genome Browser – video tutorials and training materials** https://genome.ucsc.edu/training/