
Topics
Explore selected bioinformatics resources:
- NCBI Gene and Ensembl (gene annotation databases)
- BioMart (retrieve gene list information without programming)
- NCBI GEO (experimental data repository)
- GEO2R (differential expression analysis tool)
- GO and KEGG (gene functional annotations)
- gProfiler (functional enrichment analysis tool)
Learning Objectives
By the end of this activity, you should be able to:
- Retrieve the HGNC symbol for a given human gene
- Retrieve a list of all human genes annotated with a given GO (Gene Ontology) term
- Describe the scope of a differential expression analysis
- Describe the scope of a functional enrichment analysis
- Identify differentially expressed genes in the GSE8671 dataset
- Retrieve enriched functional annotations for a gene list of interest using the gProfiler webtool
Biological databases: data types and examples
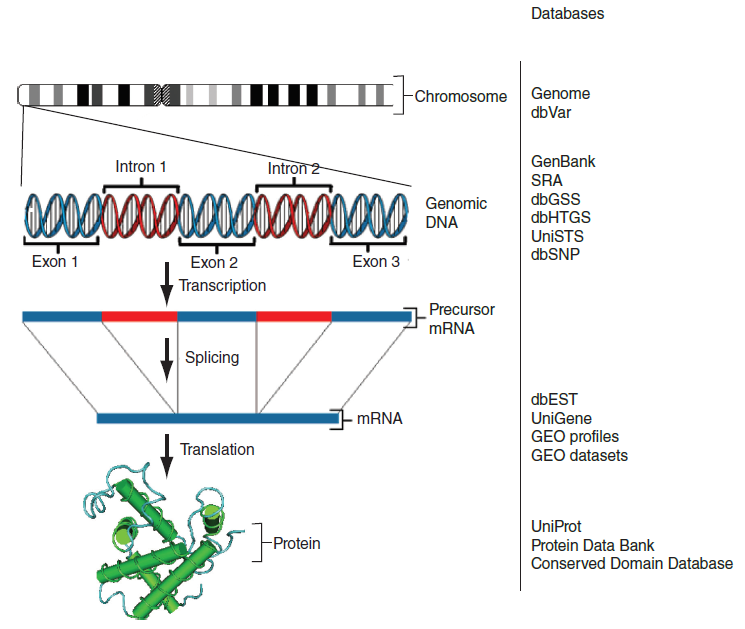
Examples of biological molecules and biological databases storing the corresponding data type (image credits: figure 2.4 from B&FG)
Gene Annotations: NCBI Gene e Ensembl
Using the two databases mentioned above (NCBI Gene and Ensembl), try to answer the following questions about the human gene ZFP36.
Note: Use this activity as an opportunity to explore the databases visually, taking your time and browsing through the different sections available for a gene of interest. Be curious and pay attention to the various types of information provided.
- On which chromosome is this gene located?
- What type of protein does it encode?
- What is its identifier (ID) in the NCBI Gene database? And in Ensembl? Are they the same?
- What is its official name (HGNC symbol)?
- How many transcripts are annotated for this gene in Ensembl? Are they all protein-coding?
- How many transcripts are annotated for this gene in NCBI Gene? Are they all protein-coding?
Take-home messages from the above activity:
- The role of identifiers in a database
- The main gene identifiers used in core gene annotation databases such as NCBI Gene, Ensembl, and, for human genes, HGNC symbols
- The meaning of MANE flags in annotated transcripts
❓ A challenge for you: What is the official gene symbol (i.e., HGNC symbol) for the NUP475 gene?
the BioMart tool @ EBI:
The Ensembl BioMart datamining tool offers a single and harmonized access point to a huge amount of information gathered together from hundreds data resources from the EBI-EMBL suite as well as from third-party databases (e.g. miRBase, HUGO, GO). It allows to perform complex queries with no need for computational or programming knowledge.
You can use BioMart to collect gene information or gene mapping among different databases in tabular format, or to get gene sequences. You can do so for all genes in a genome at once, or for just a subset of them (such as only genes on one specific region of a chromosome, or genes on one region of a chromosome associated with a given protein domain).
🎥 Short video from the Ensembl Training channel introducing BioMart
Core arguments of any BioMart search
- Mart: the combination of database (e.g. Ensembl Genes version XX) and dataset (e.g. human genes for genome build GRChYY) to be queried
- Filters: define which genes (or entries) you want to retrieve
- Values: identifiers to be used along with the filters
- Attributes: the fields/columns returned in the results table (or in sequence headers for sequence outputs)
Where to start with BioMart
- Open BioMart in a browser. (i.e., go to the Ensembl website and select “BioMart” from the top menu.)
- Follow the BioMart tutorial available among the Ensembl training materials here:
A new version of the Ensembl database is released approximately every three months, incorporating updated genome annotations. The tutorial above is based on an older Ensembl release (version 52, from 2009). While the overall structure and logic of BioMart remain consistent, some details—such as filter and attribute names—may differ in current releases. You may therefore need to adapt parts of the tutorial to match the latest version of the database.
❓ A challenge for you: How many human genes are annotated as having DNA-binding transcription factor activity (i.e., GO term GO:0003700) Find their official gene symbols (HGNC)
Analysis of Differentially Expressed Genes
Please refer to the introduction and the first challenge available on this website.
Functional Annotation of Genes: Gene Ontology (GO), KEGG and more
Gene Ontology
The Gene Ontology (GO) provides a rigorously defined set of concepts that describe the functions of gene products. It uses a controlled vocabulary to consistently describe gene functions and improve data integration, comparability, and interoperability across databases and studies. GO develops a computational model of biological systems, ranging from the molecular to the organism level, across all species in the tree of life. GO aims to provide a comprehensive representation of the current scientific knowledge about the functions of gene products, namely, proteins and non-coding RNA molecules.
The Gene Ontology (GO) provides a rigorously defined set of concepts that describe the functions of gene products. It uses a controlled vocabulary to consistently describe gene knowledge and improve data integration, comparability, and interoperability across databases and studies. GO develops a computational model of biological systems, ranging from the molecular to the organism level, across all species in the tree of life. GO aims to provide a comprehensive representation of the current scientific knowledge about the functions of gene products, namely, proteins and non-coding RNA molecules.
GO is organized into three hierarchical, structured ontologies (tree-like categories) that describe gene products in terms of molecular function, biological processe and cellular component. Specifically:
- GO Molecular Functions (MF) describe activities that occur at the molecular level, such as “DNA binding transcription factor activity” or “histone deacetylase activity”.
- Biological Processes (BP) represent the larger processes or ‘biological programs’ accomplished by multiple molecular activities. Examples of broad biological process terms are “transcription” or “signal transduction”.
- Cellular Components (CC) are the cellular structures in which a gene product performs a function, either cellular compartments (e.g., “nucleus” or “chromatin”), or stable macromolecular complexes of which they are parts (e.g., “RNA polymerase II”).
Together, annotations of a gene to terms from each of those aspects describe what specific function a gene product plays in a process and where this activity occurs in the cell.
Cellular pathways
Gene function can also be annotated within the context of cellular processes and biological pathways, providing higher-level and more informative insights, although for a smaller number of well-characterized genes compared to GO annotations.

Functional enrichment analysis of a list of genes
Functional enrichment analysis is a commonly used approach to infer relevant gene characteristics and cellular processes associated with a gene list of interest. It compares the proportion of genes associated with a given gene set in the input list to that in a background gene list, in order to determine whether the gene set is over- or under-represented. Statistical significance is typically assessed using Fisher’s exact test, a chi-squared test, or similar methods.
👉 Learn how to perferm functional enrichment analysis of a gene list of interest using the gProfiler webtool.